Our methods are offered to the scientific community as freely available resources. (Re-)distribution of the methods, in whole or in part, for commercial purposes is prohibited. CSI:FingerID and CANOPUS web services hosted by the Böcker group are for academic research and education use only. Please review the terms of service of the academic version for details. For non-academic users, the Bright Giant GmbH provides licenses and all related services. We ask that users of our tools cite the corresponding papers in any resulting publications.
Contact: 
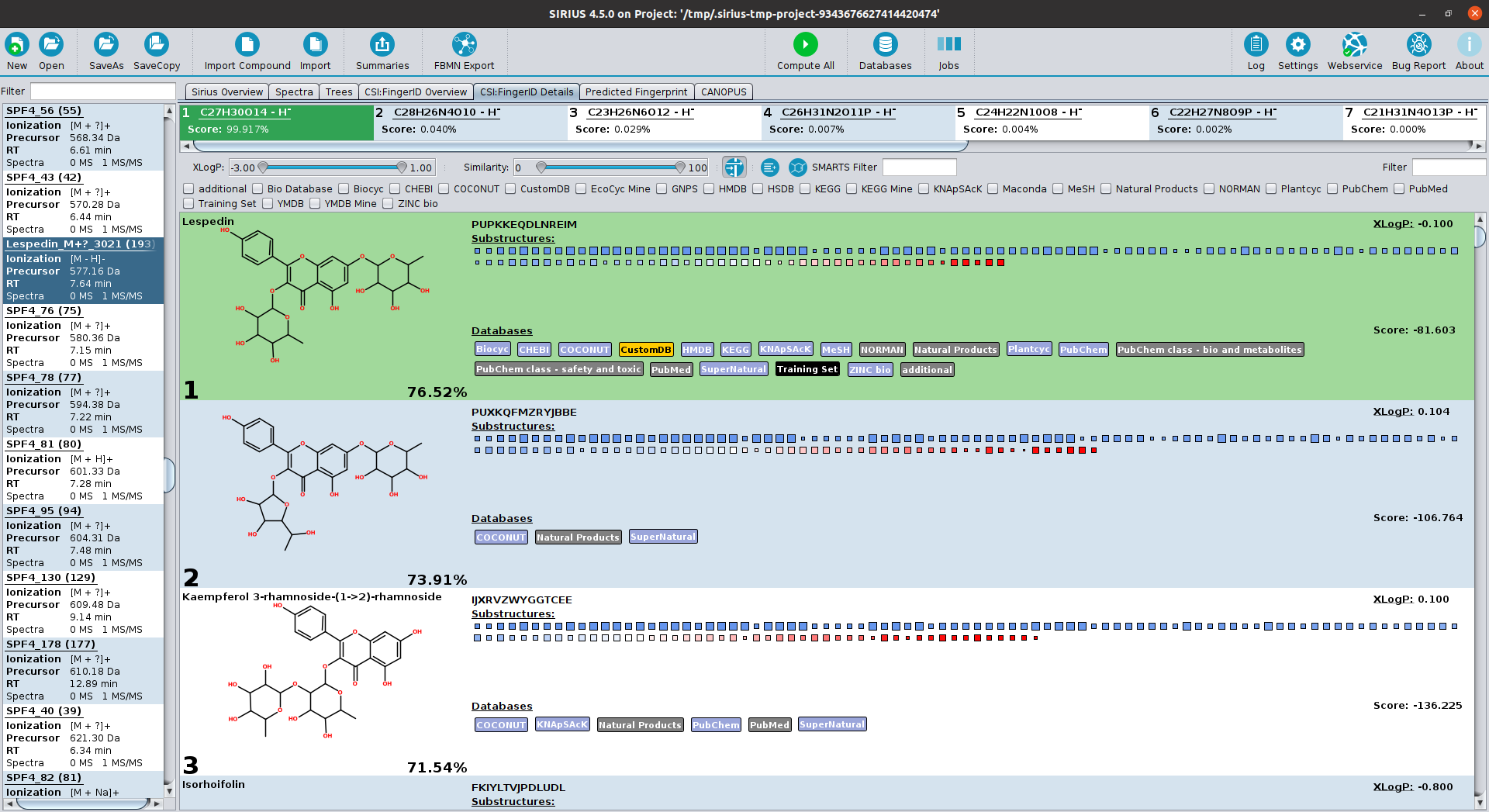
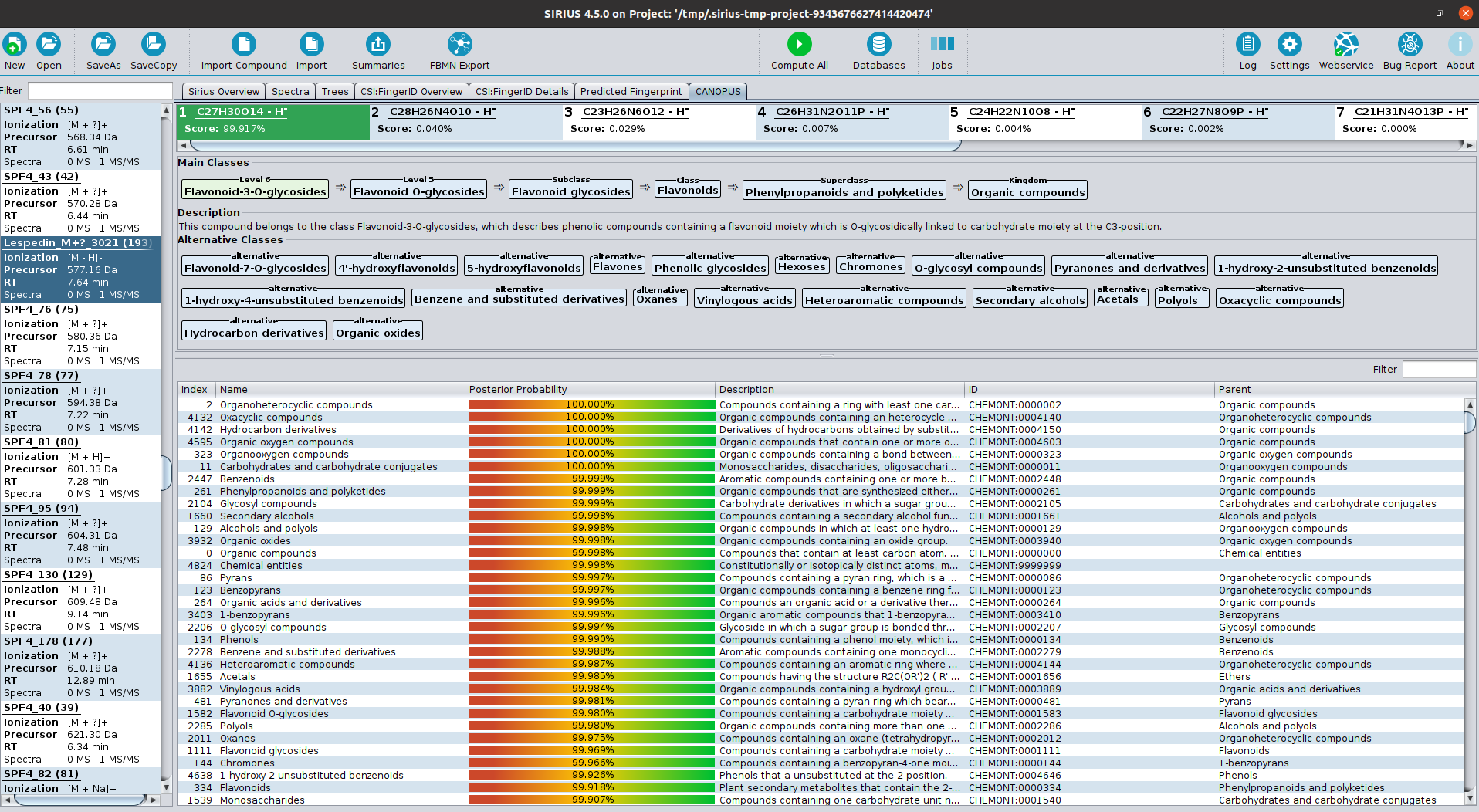
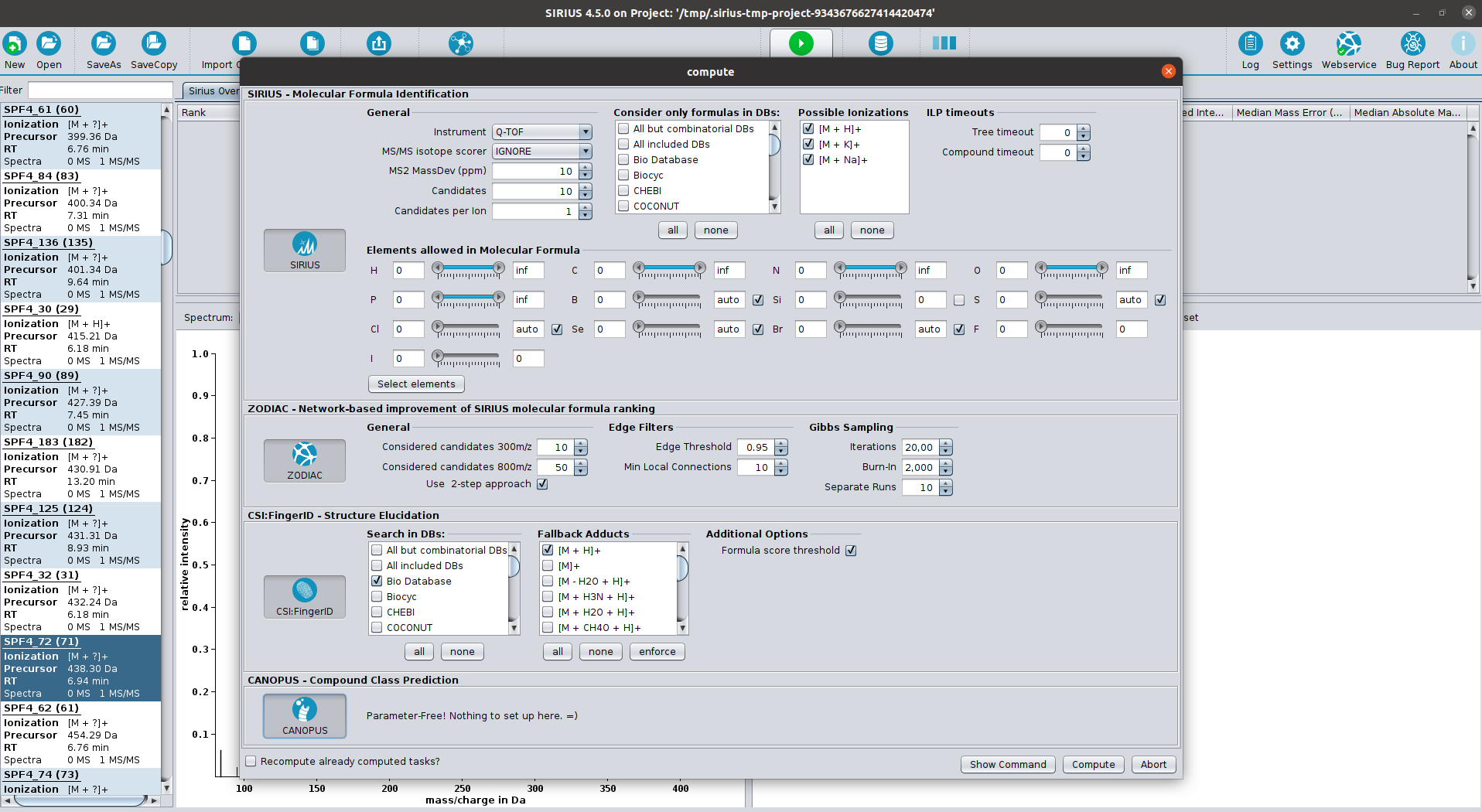
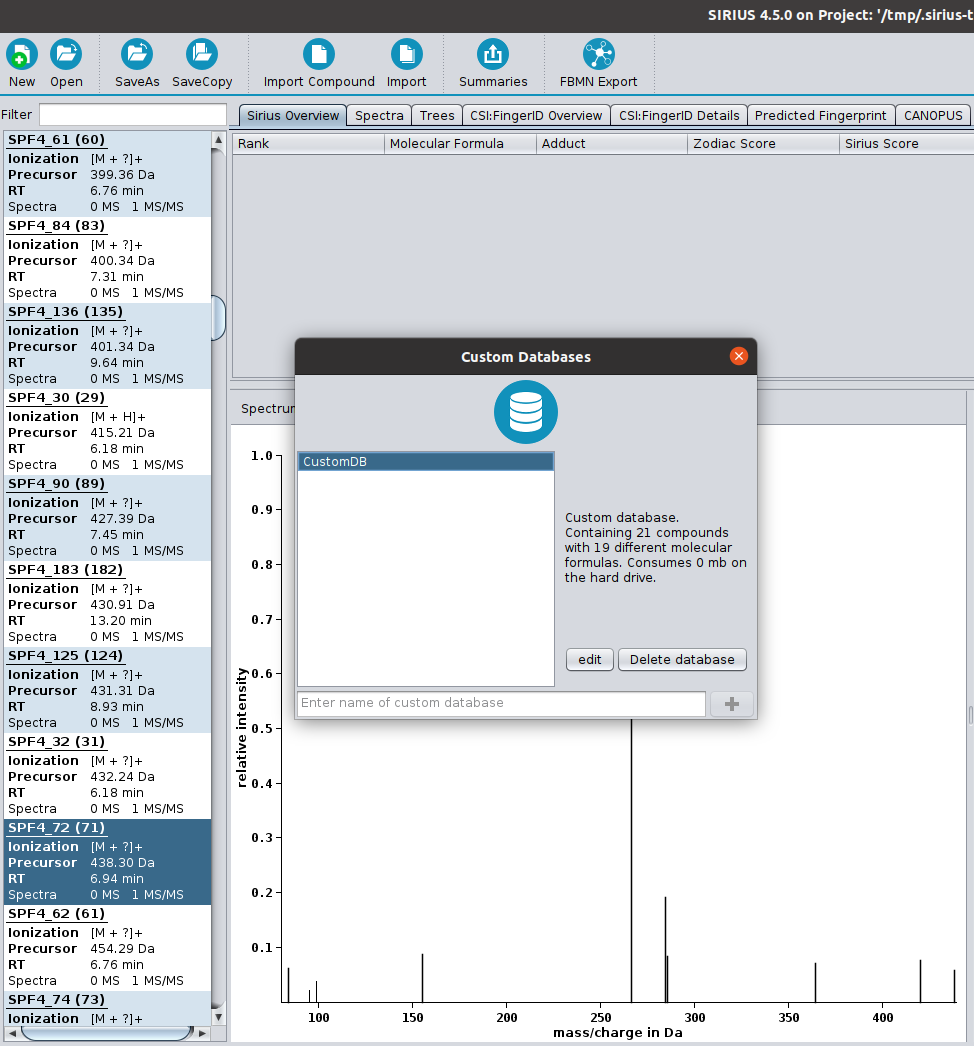
SIRIUS is a java-based software framework for the analysis of LC-MS/MS data of metabolites and other "small molecules of biological interest". SIRIUS integrates a collection of our tools, including CSI:FingerID (with COSMIC), ZODIAC and CANOPUS. In particular, both the graphical user interface and the command line version of SIRIUS seamlessly integrate the CSI:FingerID and CANOPUS web services.
Download Links
Since version 5.7.0 SIRIUS is officially available via conda (conda-forge) under the
package name sirius-ms. Native MacOS arm64 (Apple Silicon) builds are solely available
via conda.
SIRIUS+CSI:FingerID GUI and CLI - Version 5.8.6 (2024-01-27)
These versions include the Java Runtime Environment, so there is no need to install Java separately! Just download, install/unpack and execute.
- for Windows (64bit): msi / zip / msi (signed by Bright Giant)
- for Mac (64bit): pkg / zip / pkg (signed by Bright Giant)
- for Linux (64bit): zip
All (including previous) releases can be found here.
Documentation
- Online Documentation
- Video tutorials
- Bookchapter on using SIRIUS 4 (Preprint) -- does not cover the new LC-MS/MS processing option
- Demo data
- Logos for publications and presentations
Installation
The recommended method to install SIRIUS is via conda (sirius-ms) since
the package manager takes care about system library compatibility.
conda install -c conda-forge sirius-msFor Windows and MacOS, we also provide installers (msi/pkg) which should be preferred over the standalone
zip packages but might require administrator permissions for installation.
Since we do not pay Microsoft/Apple for certification, you might have to confirm that you want to trust "software from an unknown source" on Windows/MacOS when using the installers provided by the Böcker group. Therefore, we highly recommend using the signed installers provided by Bright Giant (also linked above). These installers ease the installation process by triggering no (or less) security issues of the respective OS.
See the documenntation for details.
Creating a user account
User accounts can be created directly via the SIRIUS GUI. Please, use your institutional email address. SIRIUS web services are free for academic/non-commercial use. Usually academic institutions are identified by their email domain and access will be granted automatically. In some cases, further validation of your academic/non-commercial may be required. See also SIRIUS Documentation – Account and License.
These might be especially interesting for Windows and MacOS users that have problems with the installation due to an unsigned installer (Unknown developer etc...)
Sources on GitHub
Changelog
Integration of CSI:FingerID
Fragmentation trees and spectra can be directly uploaded from SIRIUS to a CSI:FingerID web service, without the need to access the (deprecated) CSI:FingerID website. Results are retrieved from the web service and can be displayed in the SIRIUS graphical user interface. This functionality is also available for the SIRIUS command-line tool. Training structures for CSI:FingerID's predictors are available through the CSI:FingerID web API:
- https://www.csi-fingerid.uni-jena.de/v2.6/api/fingerid/trainingstructures?predictor=1 (training structures for positive ion mode)
- https://www.csi-fingerid.uni-jena.de/v2.6/api/fingerid/trainingstructures?predictor=2 (training structures for negative ion mode)
Fragmentation Tree Computation
The manual interpretation of tandem mass spectra is time-consuming and non-trivial. SIRIUS analyses the fragmentation pattern resulting in a hypothetical fragmentation tree, in which nodes are annotated with molecular formulas of the fragments and arcs (edges) represent fragmentation events (losses). SIRIUS allows for the automated and high-throughput analysis of small-compound MS data beyond elemental composition without requiring compound structures or a mass spectral database.
Isotope Pattern Analysis
SIRIUS deduces molecular formulas of small compounds by ranking isotope patterns from mass spectra of high resolution. After preprocessing, the output of a mass spectrometer is a list of peaks which corresponds to the masses of the sample molecules and their abundance. In principle, elemental compositions of small molecules can be identified using only accurate masses. However, even with very high mass accuracy, many formulas are obtained in higher mass regions. High resolution mass spectrometry allows us to determine the isotope pattern of sample molecule with outstanding accuracy and apply this information to identify the elemental composition of the sample molecule. SIRIUS can be downloaded either as graphical user interface (see Sirius GUI) or as command-line tool.
Main citations
Kai Dührkop, Markus Fleischauer, Marcus Ludwig, Alexander A. Aksenov, Alexey V. Melnik, Marvin Meusel, Pieter C. Dorrestein, Juho Rousu, and Sebastian Böcker, SIRIUS 4: Turning tandem mass spectra into metabolite structure information. Nature Methods 16, 299–302, 2019.
Martin A. Hoffmann and Louis-Félix Nothias and Marcus Ludwig and Markus Fleischauer and Emily C. Gentry and Michael Witting and Pieter C. Dorrestein and Kai Dührkop and Sebastian Böcker Assigning confidence to structural annotations from mass spectra with COSMIC bioRxiv, 2021. (Cite if you are using: CSI:FingerID, COSMIC)
Kai Dührkop, Louis-Félix Nothias, Markus Fleischauer, Raphael Reher, Marcus Ludwig, Martin A. Hoffmann, Daniel Petras, William H. Gerwick, Juho Rousu, Pieter C. Dorrestein and Sebastian Böcker. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nature Biotechnology, 2020. (Cite if you are using CANOPUS)
Yannick Djoumbou Feunang, Roman Eisner, Craig Knox, Leonid Chepelev, Janna Hastings, Gareth Owen, Eoin Fahy, Christoph Steinbeck, Shankar Subramanian, Evan Bolton, Russell Greiner, David S. Wishart. ClassyFire: automated chemical classification with a comprehensive, computable taxonomy. Journal of Cheminformatics 8, 61, 2016. (ClassyFire publication; cite this if you are using CANOPUS)
Marcus Ludwig, Louis-Félix Nothias, Kai Dührkop, Irina Koester, Markus Fleischauer, Martin A. Hoffmann, Daniel Petras, Fernando Vargas, Mustafa Morsy, Lihini Aluwihare, Pieter C. Dorrestein, Sebastian Böcker. Database-independent molecular formula annotation using Gibbs sampling through ZODIAC. Nature Machine Intelligence 2, 629–641, 2020. (Cite if you are using ZODIAC)
Kai Dührkop and Sebastian Böcker. Fragmentation trees reloaded. Journal of Cheminformatics 8, 5, 2016. (Cite this for fragmentation pattern analysis and fragmentation tree computation)
Kai Dührkop, Huibin Shen, Marvin Meusel, Juho Rousu, and Sebastian Böcker. Searching molecular structure databases with tandem mass spectra using CSI:FingerID. Proceedings of the National Academy of Sciences U S A 112(41), 12580-12585, 2015. (cite this when using CSI:FingerID)
Sebastian Böcker, Matthias C. Letzel, Zsuzsanna Lipták and Anton Pervukhin. SIRIUS: decomposing isotope patterns for metabolite identification. Bioinformatics 25(2), 218-224, 2009. (Cite this for isotope pattern analysis)
Additional citations
Marcus Ludwig, Kai Dührkop and Sebastian and Böcker. Bayesian networks for mass spectrometric metabolite identification via molecular fingerprints. Bioinformatics, 34(13): i333-i340. 2018. Proc. of Intelligent Systems for Molecular Biology (ISMB 2018). (Cite for CSI:FingerID Scoring)
W. Timothy J. White, Stephan Beyer, Kai Dührkop, Markus Chimani and Sebastian Böcker. Speedy Colorful Subtrees. In Proc. of Computing and Combinatorics Conference (COCOON 2015), volume 9198 of Lect Notes Comput Sci, pages 310-322. Springer, Berlin, 2015. (cite this on why computations are swift, even on a laptop computer)
Huibin Shen, Kai Dührkop, Sebastian Böcker and Juho Rousu. Metabolite Identification through Multiple Kernel Learning on Fragmentation Trees. Bioinformatics, 30(12):i157-i164, 2014. Proc. of Intelligent Systems for Molecular Biology (ISMB 2014). (Introduces the machinery behind CSI:FingerID)
Imran Rauf, Florian Rasche, François Nicolas and Sebastian Böcker. Finding Maximum Colorful Subtrees in practice. J Comput Biol, 20(4):1-11, 2013. (More, earlier work on why computations are swift today)
Heinonen, M.; Shen, H.; Zamboni, N.; Rousu, J. Metabolite identification and molecular fingerprint prediction through machine learning. Bioinformatics, 2012. Vol. 28, nro 18, pp. 2333-2341. (Introduces the idea of predicting molecular fingerprints from tandem MS data)
Florian Rasche, Aleš Svatoš, Ravi Kumar Maddula, Christoph Böttcher, and Sebastian Böcker. Computing Fragmentation Trees from Tandem Mass Spectrometry Data. Analytical Chemistry (2011) 83 (4): 1243–1251. (Cite this for introduction of fragmentation trees as used by SIRIUS)
Sebastian Böcker and Florian Rasche. Towards de novo identification of metabolites by analyzing tandem mass spectra. Bioinformatics (2008) 24 (16): i49-i55. (The very first paper to mention fragmentation trees as used by SIRIUS)
License
Starting with version 4.4.27, SIRIUS is licensed under the GNU Affero General Public License (GPL). If you integrate SIRIUS into other software, we strongly encourage you to make the usage of SIRIUS as well as the literature to cite transparent to the user.

