Our article “Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra” has just appeared in Nature Biotechnology. Congrats to Kai and all co-authors!
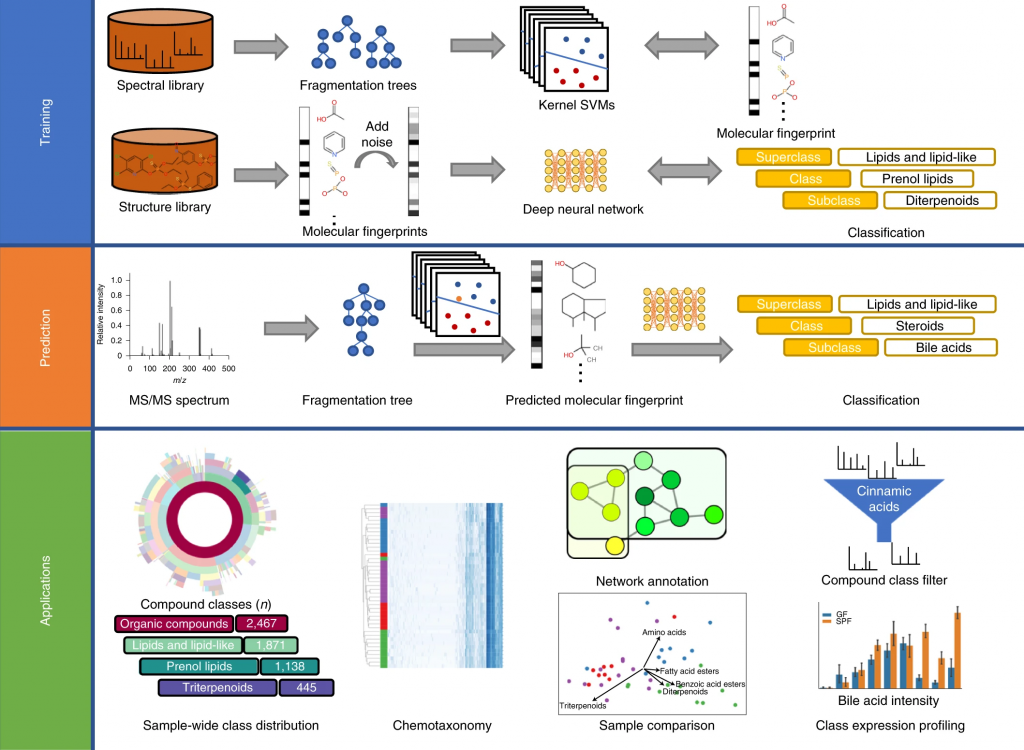
In short: CANOPUS is a computational tool for systematic compound class annotation. It uses a deep neural network to predict 2,497 compound classes from fragmentation spectra, including all biologically relevant classes. From the machine learning perspective, the interesting part is that different levels of the neural network are trained using different data (heterogeneous training). CANOPUS explicitly targets compounds for which neither spectral nor structural reference data are available, and even predicts classes completely lacking tandem mass spectrometry training data. In evaluation using reference data, CANOPUS reached very high prediction performance (average accuracy of 99.7% in cross-validation) and outperformed four (rather advanced) baseline methods. We used CANOPUS to investigating the effect of microbial colonization in the mouse digestive system, for analyzing the chemodiversity of different Euphorbia plants, and for the structural elucidation of a novel marine natural product.
CANOPUS is already available to users through SIRIUS 4.5, which was released last Thursday. See also the designated CANOPUS page. A view-only version of the article is available here.
Full citation: K. Dührkop, L.-F. Nothias, M. Fleischauer, R. Reher, M. Ludwig, M. A. Hoffmann, D. Petras, W. H. Gerwick, J. Rousu, P. C. Dorrestein, and S. Böcker. Systematic classification of unknown metabolites using high-resolution fragmentation mass spectra. Nat Biotechnol, 2020. https://doi.org/10.1038/s41587-020-0740-8