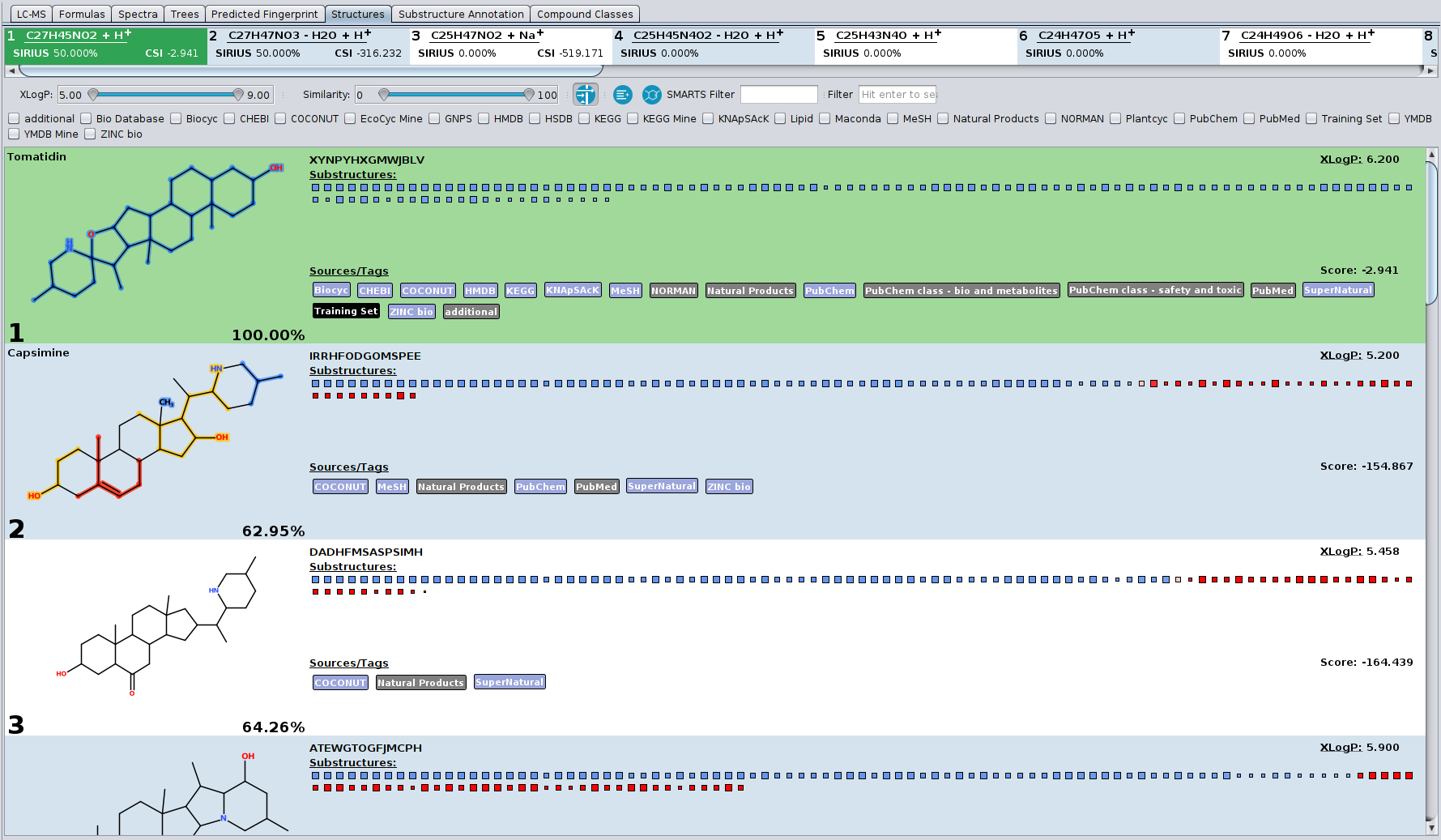
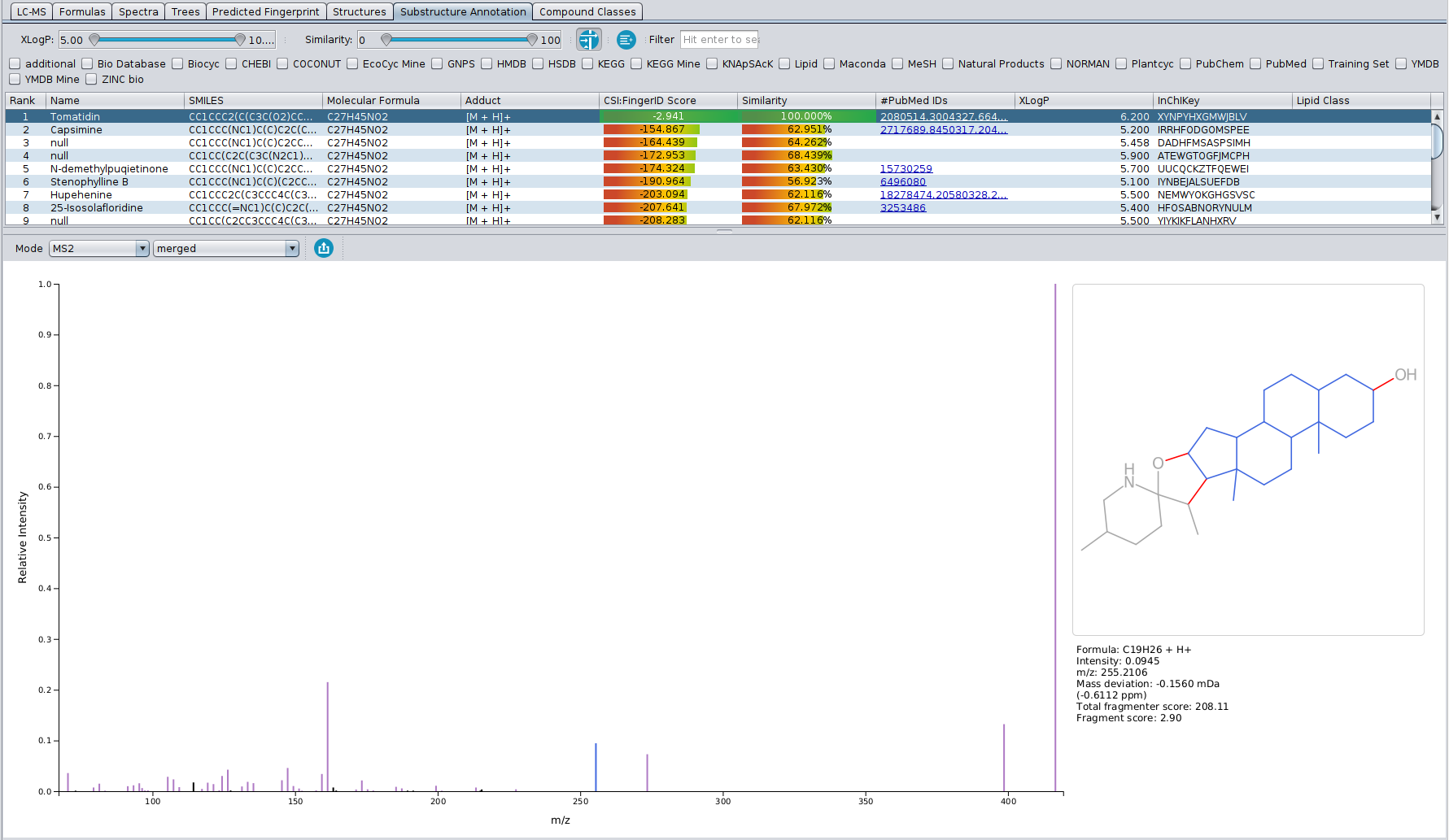
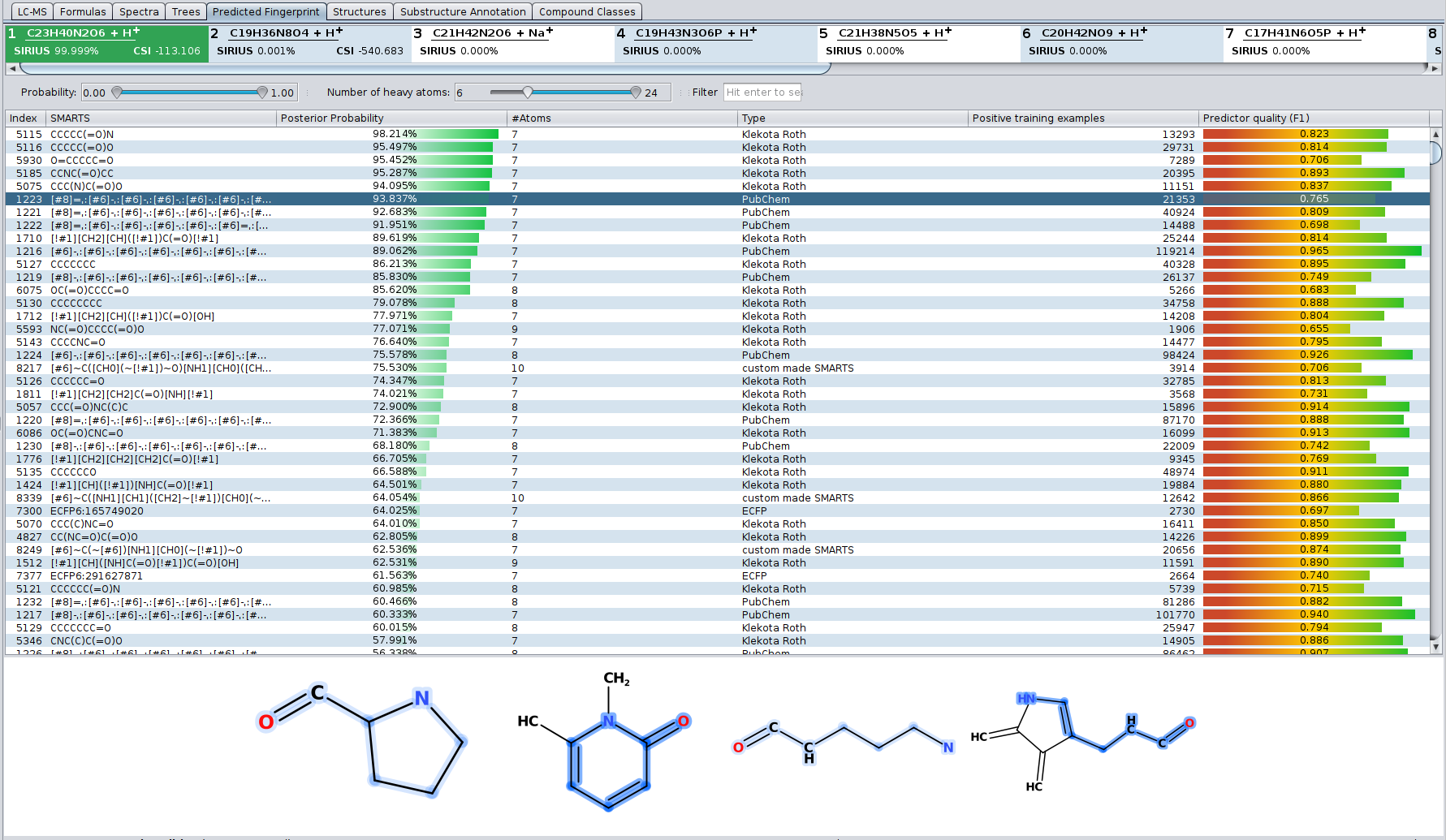
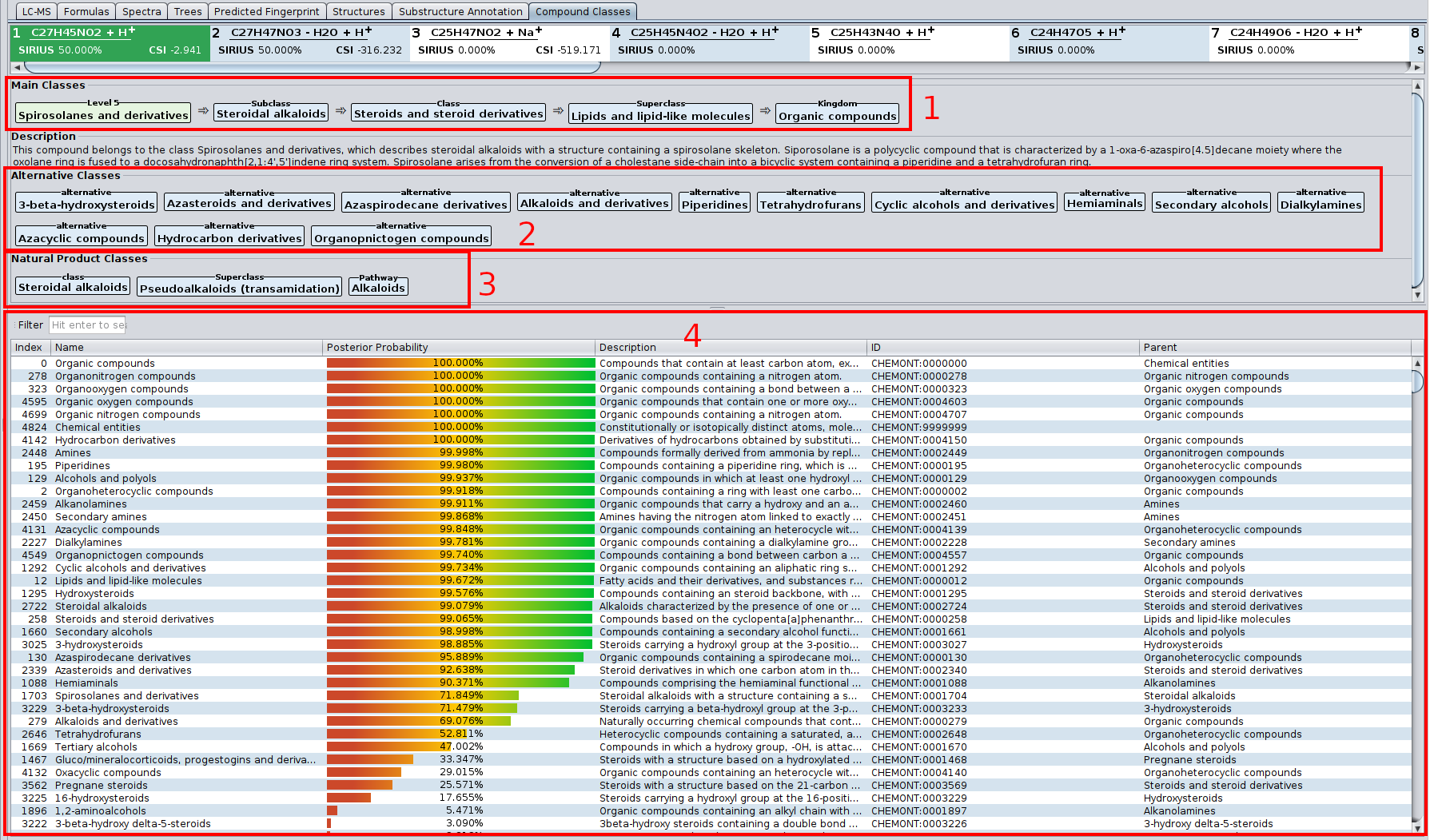
Our article “High-confidence structural annotation of metabolites absent from spectral libraries” has just appeared in Nature Biotechnology. Congrats to Martin and all co-authors!
In short, COSMIC allows you to assign confidence to structure annotations. For every structure annotated by CSI:FingerID, COSMIC provides a confidence score (a number between 0 and 1) that tells you how likely it is that this annotation is correct. This is similar in spirit to what is done in spectral library search: Not only is the cosine score used to decide which candidate best fits to the query spectrum; in addition, we use the cosine of the top-scoring candidate (the hit) to decide whether it is likely correct (say, above 0.8), incorrect (say, below 0.6) or in the “twilight” in-between. If you have been using CSI:FingerID for some time, you might have noticed that finding such thresholds is not possible for the CSI:FingerID score. COSMIC closes this gap and tells you if an annotation is likely correct or incorrect.
Doing so is undoubtedly convenient in practice; but this is not what COSMIC is all about. What we can do now is to sort all hits in a dataset or even a repository with respect to confidence, and then concentrate our downstream analysis on high-confidence annotations. Next, we can replace the “usual” structure databases we search in by a structure database made entirely from hypothetical structures generated by combinatorics, machine learning or in silico enzymatic reactions.
We demonstrate COSMIC’s power by generating a database of hypothetical bile acid structures, combinatorially adding amino acids to bile acid cores, yielding 28,630 plausible bile acid conjugate structures. We then searched query MS/MS data from a mice fecal dataset in this combinatorial database, and used the COSMIC confidence score to distinguish between hits that are likely correct or incorrect. We manually evaluated the top 12 hits and found that 11 annotation (91.6%) were likely correct; two annotations were further confirmed using synthetic standards. All 11 bile acid conjugates are “truly novel”, meaning that we could not find those structures in PubChem or any other structure database (or publication). Whereas reporting 11 novel bile acid conjugates may appear rather cool, we argue it is even cooler that we did this without a biological hypothesis beyond “there might be bile acid conjugates out there which nobody knows about”; and that COSMIC found the top bile acid conjugate annotations in a fully automated manner and in in a matter of hours.
We have also annotated 2,666 LC-MS/MS runs from human samples with molecular structures which are currently absent from HMDB, and for which no MS/MS reference data are available; and finally, 17,414 LC-MS/MS runs with annotations for which no MS/MS reference data are available. We hope that some of them might be of interest to you.
If you have an idea of hypothetical structures, similar to the bile acid conjugates, to be searched against thousands of datasets, please let us know.
COSMIC’s confidence score is available through SIRIUS since version 4.8, download here.