We are happy to introduce CANOPUS, a tool for the comprehensive annotation of compound classes from MS/MS data (certain restrictions apply, see below). In principle, CANOPUS is doing something similar as CSI:FingerID: Whereas CSI:FingerID can tell you what substructures are part of the query compound, CANOPUS does so for compound classes. The differences between both tasks are subtle but have massive consequences. See this preprint on the details of this difference, how CANOPUS works, how good it works etc.
At present, CANOPUS predicts 1270 compound classes. In more detail, CANOPUS predicts ClassyFire compound classes. ClassyFire is not the first but, to the best of our knowledge, by far the most comprehensive approach to assign classes solely from structure. (This last point is key, as this allows us to assign thousands of classes for millions of molecular structures.) Please have a look there if you use CANOPUS: Certain compound class definitions may be not what you expect. For example, we found that many phytosteroids are classified as bile acids in ClassyFire. While the biochemical origin of both classes is very different, they are structural very similar and, therefore, represented by the same class in the ClassyFire ontology.
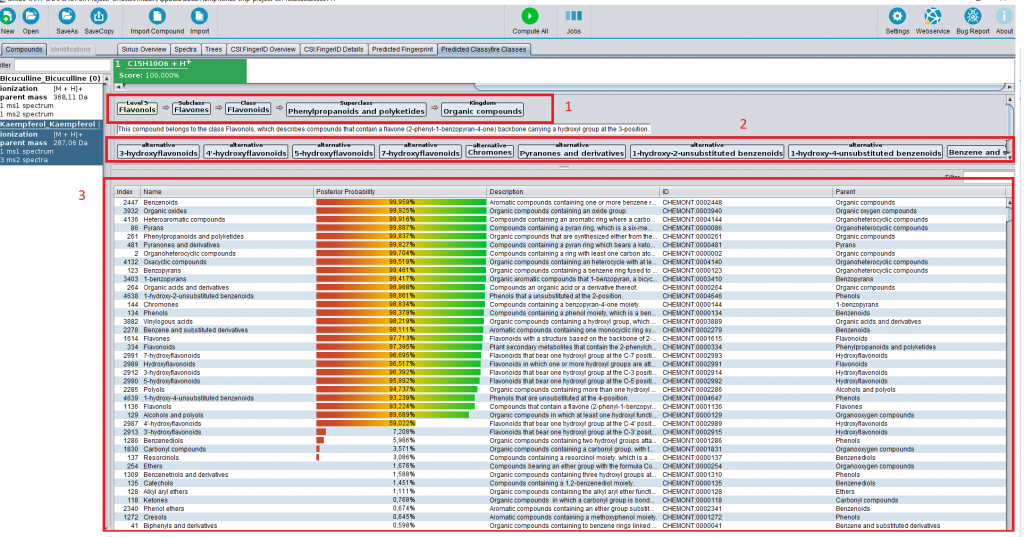
You can download, install and use CANOPUS through SIRIUS 4.4. You will notice a new tab where you can access, for each compound, all compound classes it does or does not belong to (and, how sure we are about that). Fancier visualizations (see the preprint) will be made available with upcoming releases.
ps. Clearly, CANOPUS is comprehensive only within the limits of the LC-MS/MS technology: If a compound does not ionize, if no fragmentation spectrum is recorded in Data Dependent Acquisition, if a compound does not show any fragmentation, if multiple compounds are fragmented in a single spectrum etc, then CANOPUS cannot help you. We don’t do magic. Also, CANOPUS is limited by the available (structure and MS/MS) training data; but several years of thinking have been invested to get the most out of it.